To import mass spec data into iolite, a few key pieces of information are required. The most basic of these are the time the data were collected and what masses were measured. Without this information, iolite can’t import the data and represent it in time series, and time is very important when you’re dealing with time-dependent phenomena such as mass bias etc. Most mass spectrometers produce files that contain this information, however the Nu Plasma .run files are a little trickier. They don’t contain the names of the masses collected (i.e. the channel names). There are a couple of ways around this, such as decoding the associated .nrf file (more on .nrf files below), and we have implemented these in previous versions of iolite. However for iolite v4, we decided that (all things considered) it would be easiest to write individual importers for .run files.
Here at UniMelb, we have two Nu Plasma multi-collector mass specs, and both produce .run files. We do create a variety of .run files, but getting the data into iolite is mostly straightforward although it does require editing a bit of code. If the phrase “editing the code” makes you nervous, don’t worry: it’s pretty hard to actually break anything and this note is here to guide you (there are just three minor edits that you need to make). If you do have any issues, you can always post on the iolite forum for help.
A quick look at the .run file format
Firstly, lets take a look inside a Nu Plasma .run file. If you open a .run file in a simple text editor (not Word) such as Notepad++ or Atom (both free), you can see that the first line contains the version information about the file. This is then followed by a bunch of lines with either a few numbers or some short text on it. This is the header information (usually just referred to as the “headers”), and we can mostly ignore it here. It does have information about which collectors were enabled etc, but there’s not enough info here to import our data into iolite.
If you scan down, you’ll see a line that just has the words “Spare Text” on it, usually around Line 46. This line marks the start of the actual data collected by the mass spec (and the end of the headers). Just above this Spare Text line is some information about the run, including the run start time, and the sample name entered when it was run. It also has which .nrf file was used to collect the data. A .nrf file is just an instructions file that tells the mass spec what masses to measure, how long to measure zeroes etc. In previous versions of iolite, we would decode that .nrf file to get the channel names. However, if your mass spec has a non-standard collector array (which is semi-common) we can’t determine which masses were measured from the .nrf file, and everything breaks down… which is why we’re editing some code here. Take note of the name of the .nrf file recorded in your file. We’ll use that infomation later.
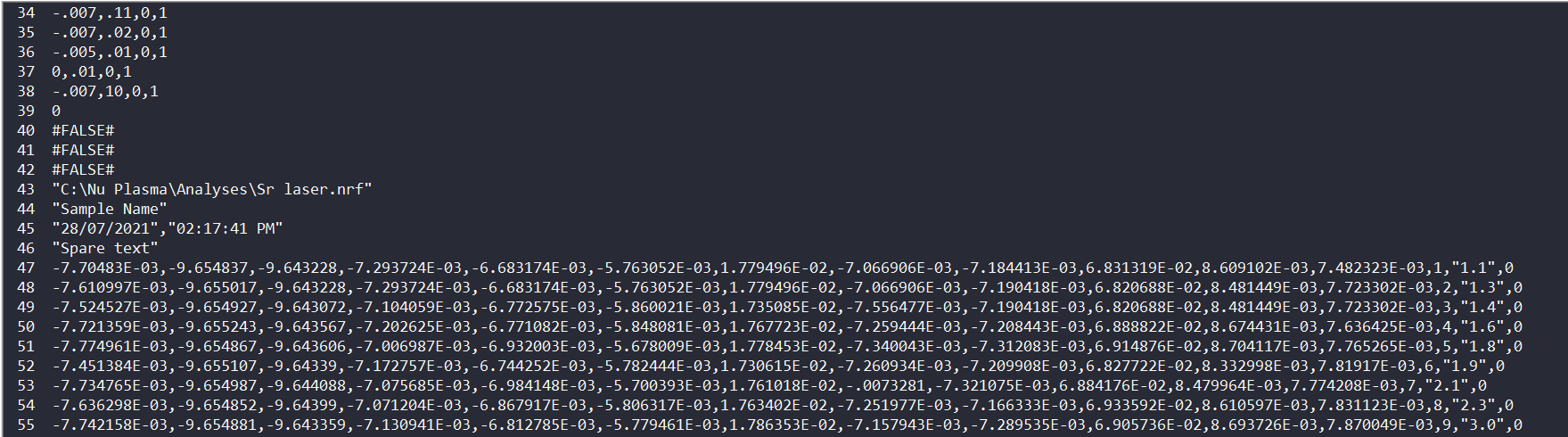
Below the Spare Text line are lines of comma-separated data. If you were to open this file in something like Excel, each value between the commas would be put into its own column. The number of columns depends on your setup, but the last three columns are usually a simple number that increases by 1 on each line; a quoted number; and a 0 or a 1. The first of these last three columns is just the measurement number (which is why it’s incremented by 1 on each line). The second of these (the quoted number) is the timestamp for this measurement. If, for example, your mass spec is recording a measurement every 0.2 seconds the values in this column will increase by 0.2 on each line (there may be a second or so setup time). The final column represents whether this measurement was considered a zero or sample measurement, but we can ignore this because we’ll create adjustable selections in iolite to determine what was zeros and sample after import.
Now to the actual data: each column represents a collector, and will become a channel in iolite. The channels start with the highest mass on the left. The mass difference between each column depends (again) on your setup. Naming these columns is what we’ll be editing in our importer.
Creating your own importer
An importer in iolite v4 is just a python script. You can have as many of these as you like, and you just need to tell iolite where you’re storing them so that iolite will use them when you try to import a file. You can set where your importers are stored in iolite’s Preferences, in the Paths section, labelled Importers. By default, importers are stored in your iolite folder, which is normally in your Documents folder unless you chose a different location when you installed iolite. If you click on the -> button at the end of the Importers Path, it’ll take you to the folder where your importers are stored. This folder might be empty: that’s OK. You’re about to put a script in this folder.
I would recommend that you download either Notepad++ or Atom so that you can edit your script. You can’t do this in something like Word. Once you have your text editor installed, you can get the code. I’ve already written a basic importer script for .run files that does almost everything. You can just download this script and edit it to accept your .run files. You can download the file from here. Make sure that you click on the Raw button to see the raw code (rather than the webpage). Once you’ve got the raw code, copy all of it and paste it into a new blank document in your text editor. Then save it as something like “Sr_run_importer.py”. Make sure that the file extension is “.py” so that iolite will know that this is a python script. Now we can edit it to make it work for your files.
Editing the importer
There are just three areas that you need to adjust for your file. The first is the metadata at the top of the script. When iolite reads the script, it needs to know what type of function this script performs (e.g. import data, DRS, QAQC module, etc). You just need to change the Name, Authors, and Description; the rest can remain the same. The Name is the only important one of these three, and should short but descriptive.
The next part to edit is the correct_format() function. When you select a file to import into iolite, iolite runs this function in each importer to see if the importer can load the file. The function searches the content of a file to see if it has something that identifies it as a file this particular importer can open and returns True (i.e. it can load this file) if it finds the text. In this case, we’ll search for the .nrf line just above the Spare Text line described above. This line will be in every file and won’t change (unlike the other metadata etc).
In this case, the .nrf file in our example was “Sr laser.nrf”. We put this into the line starting with nrf_regex(the line underlined in yellow below; don’t worry if you editor doesn’t show the vertical lines like mine, these are just visual guides). We put a backslash before the ‘.’ but otherwise it matches what was in our file’s headers, and only includes the file name, not the full file path. Make sure that you get the uppercase and lowercase letters correct (the search is case sensitive.)
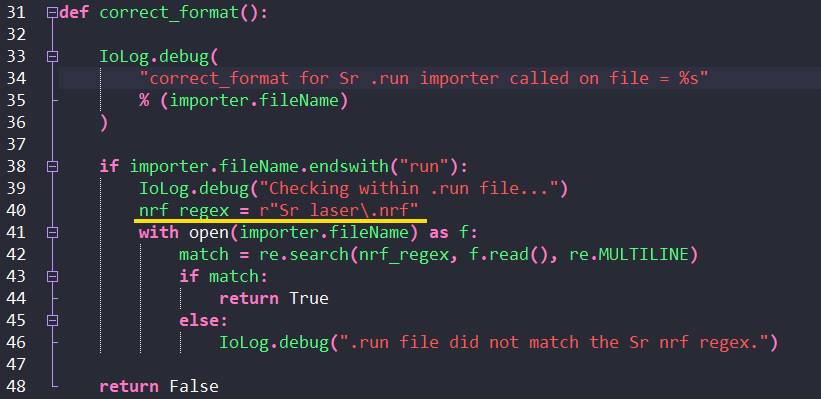
For extra points you can edit the debug message (Line 34 in our example) to say which importer was checking the file. This line is just there to help track whether the importer ran, so it’s not essential to change this, but it can help track down issues later. There are two other debug lines on Lines 46 and 55 that you can also change if you like.
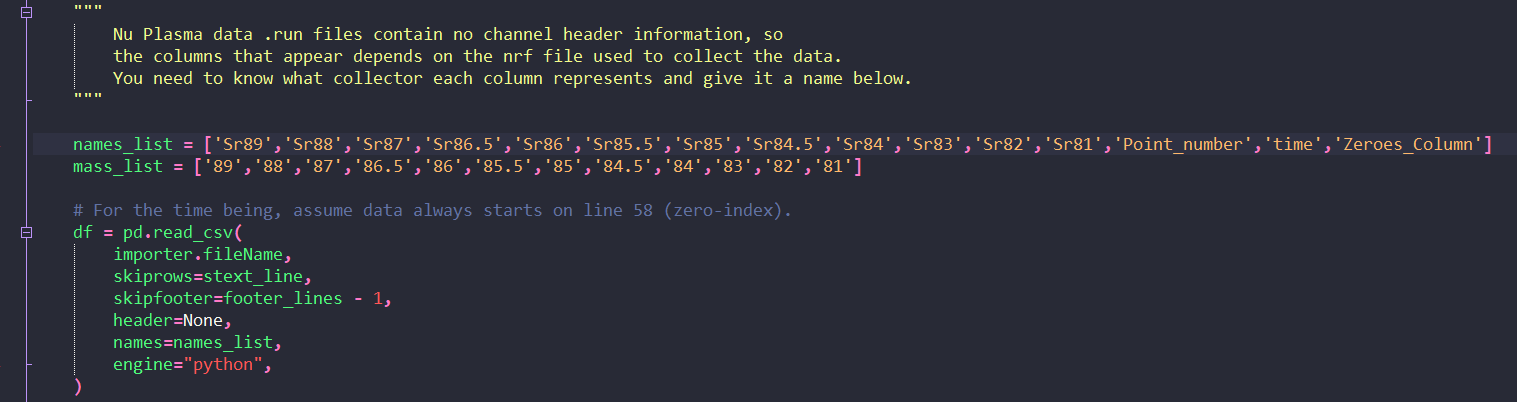
And now for the important part: to name the channels in your file. On Lines 111 and 112, you’ll see two lists: a names_list and a masses_list. Each one is just a comma-separated list of names or numbers. The names_list is what each channel in your file should be called. You just need to update this list with your masses making sure that each channel’s name is enclosed in single quotation marks, and is followed by a comma. This list also has the point number, measurement time and zeros column that I described above. Make sure you leave these in. Then edit the masses_list: it is in the same order as the names_list, but you don’t need to include the point number, measurement time or zeroes column in this list. Make sure that each value is enclosed by single quotation marks.
Save your file and that’s it: you’re done. You can now try it out in iolite.
Using the importer
For iolite to recognise the new importer, you’ll need to restart iolite. Once iolite has re-started, you should be able to see your new importer in the Plugin Manager. If you go to Tools -> Plugins -> Manage, iolite will show you a list of plugins that have been loaded in a list of the left hand side of the window. There is a search bar at the top of this list. Start typing the Name of your importer into this list, and it should appear and have a green light next to it if it was loaded properly. If it doesn’t appear, make sure that the file you’ve been working on is in the folder listed in Preferences -> Paths -> Importers.
If your importer appears and is green-lit, you can try it out by clicking the Import file button as you normally would to import files into iolite. Select your .run file and it should import.
If it doesn’t import correctly, you can check your debug messages in the Message View and contact us in the iolite forum for more help. If you have any other questions about loading .run files, please feel free to post here.