Happy new year and welcome to the first release notes highlights note!
We get the feeling that new features or changes often go unnoticed in our release notes that are displayed when updating. Since we don’t want people to miss out on the latest features, we thought we would start posting the release notes with graphical highlights, more details, and corresponding discussions on our forum.
Highlights
Viewing limits of detection
The limits of detection (LOD) have been calculated by iolite according to the following three publications for quite some time:
Originally, this calculation was done by the trace elements data reduction scheme (DRS). So, if a selection was changed after running the DRS, to get updated LODs, the DRS would need to run again. To get around this limitation, we’ve started storing the sensitivity around a selection (as a selection property) when the DRS runs and calculating LODs on demand. This meant that the LODs could be updated any time a selection was adjusted assuming that the local sensitivity has not changed significantly.
We also did not have a way to visualize the LODs previously. They could be shown in the results data table or a custom python script could be used to plot them, but there was no built-in way to see how good they were or, perhaps more importantly, check if anything unusual was going on. To remedy this, we’ve added a control to the results summary stats area that allows you to select a LOD type to include below the summary stats plot as shown below.
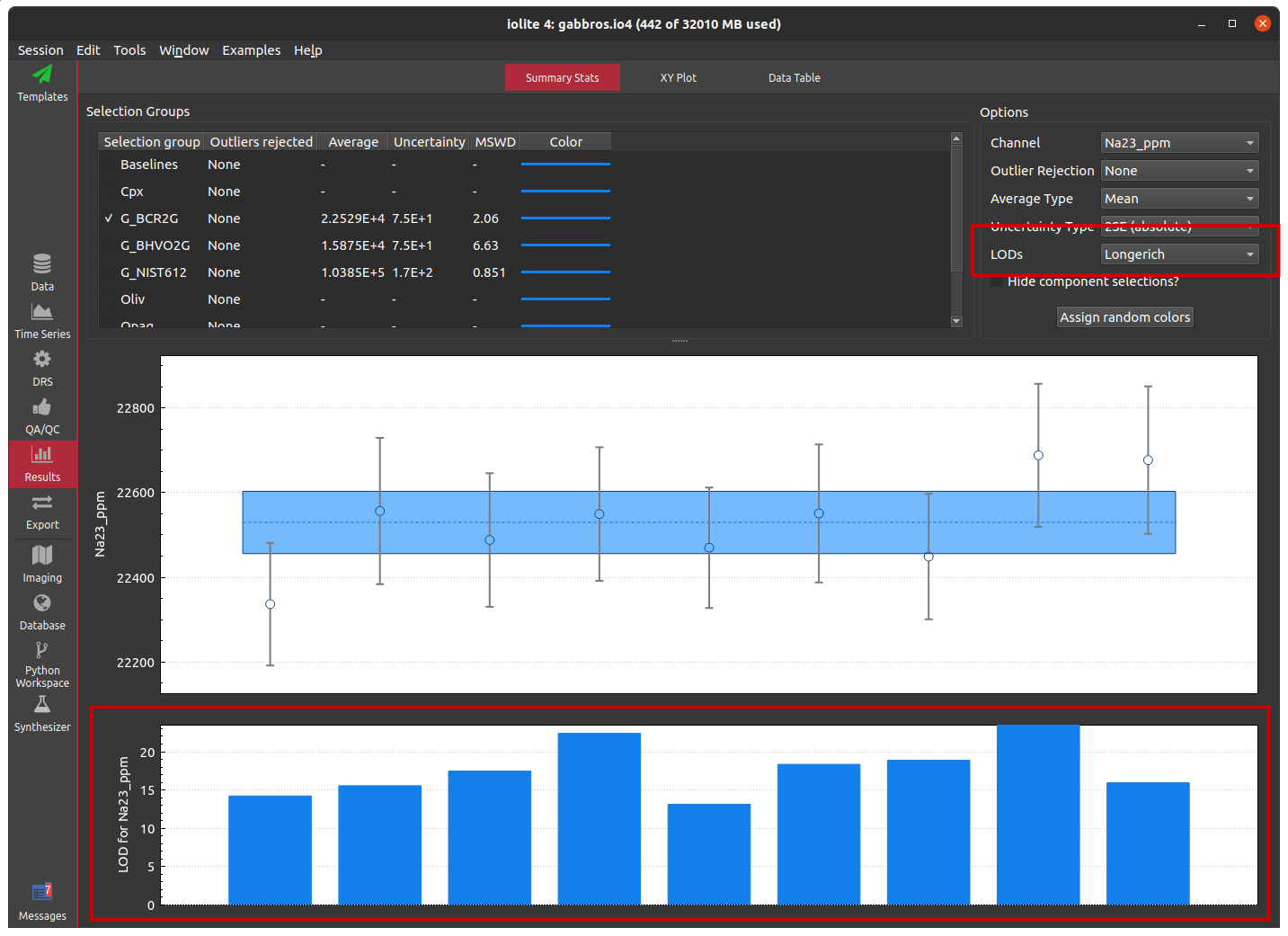
Zip importer
Some instruments produce a lot of files, and some users prefer to split each analysis into its own file. The result can be hundreds or thousands of files representing a single day of work. This can be inconvenient, but it can also consume a lot of disk space. Rather than keeping all of these files separate, it is now possible to zip them into one archive and have iolite read the data from the zip. This is great because text-based data files can be highly compressed saving tons of space, and it also makes it more convenient by having only one file to keep track of and move around. Behind the scenes, iolite extracts the zip to a temporary folder that is automatically deleted and uses the folder importer to load whatever data are found in the zip. So, it should work with any data format supported by iolite.
Zooming in external image alignment
Depending on the resolution of your images (external or internal), it was sometimes difficult to align images accurately because we did not have an option to zoom into the images to better select or align points of interest. We have addressed this by adding some zoom controls to the bottom right corner of the alignment window. When zoomed in, additional scroll bars will appear to help you navigate the area. With the image zoomed in you can check the “.coord” file alignment, manually transform the external image onto your laser images, or select points for a 3 point transformation.
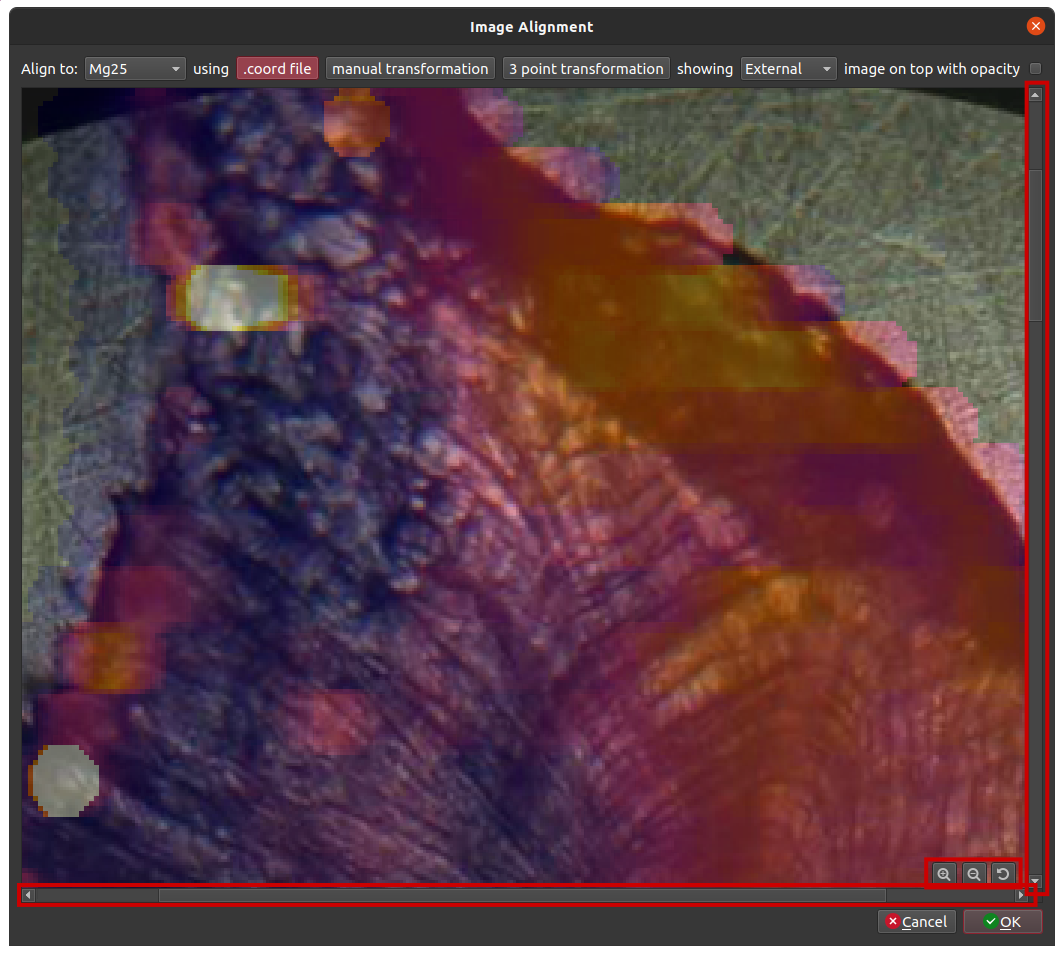
Percentile auto scaling for imaging
Manually adjusting the color scale limits to get them just right in imaging can be very satisfying. However, for users acquiring dozens (or hundreds!) of analytes and/or images, this process can be very tedious, and that satisfaction is replaced with exhaustion and frustration. Earlier versions of iolite had a few options to help in this regard: auto scaling to the minimum/maximum or the mean ± 2 or 3 standard deviations. These options work well in some scenarios, but in cases involving outliers neither of them produce useful results.
New in this release are a few options for percentile-based auto scaling (initiated by the forum post here. The options are for 90th, 95th, 99th, 99.9th percentile. When used for the lower limit, these options put the lower 10, 5, 1, or .1 percent of the data below the lowest color. Similarly, when used for the upper limit, these options put 10, 5, 1, or .1 percent of the data above the highest color.
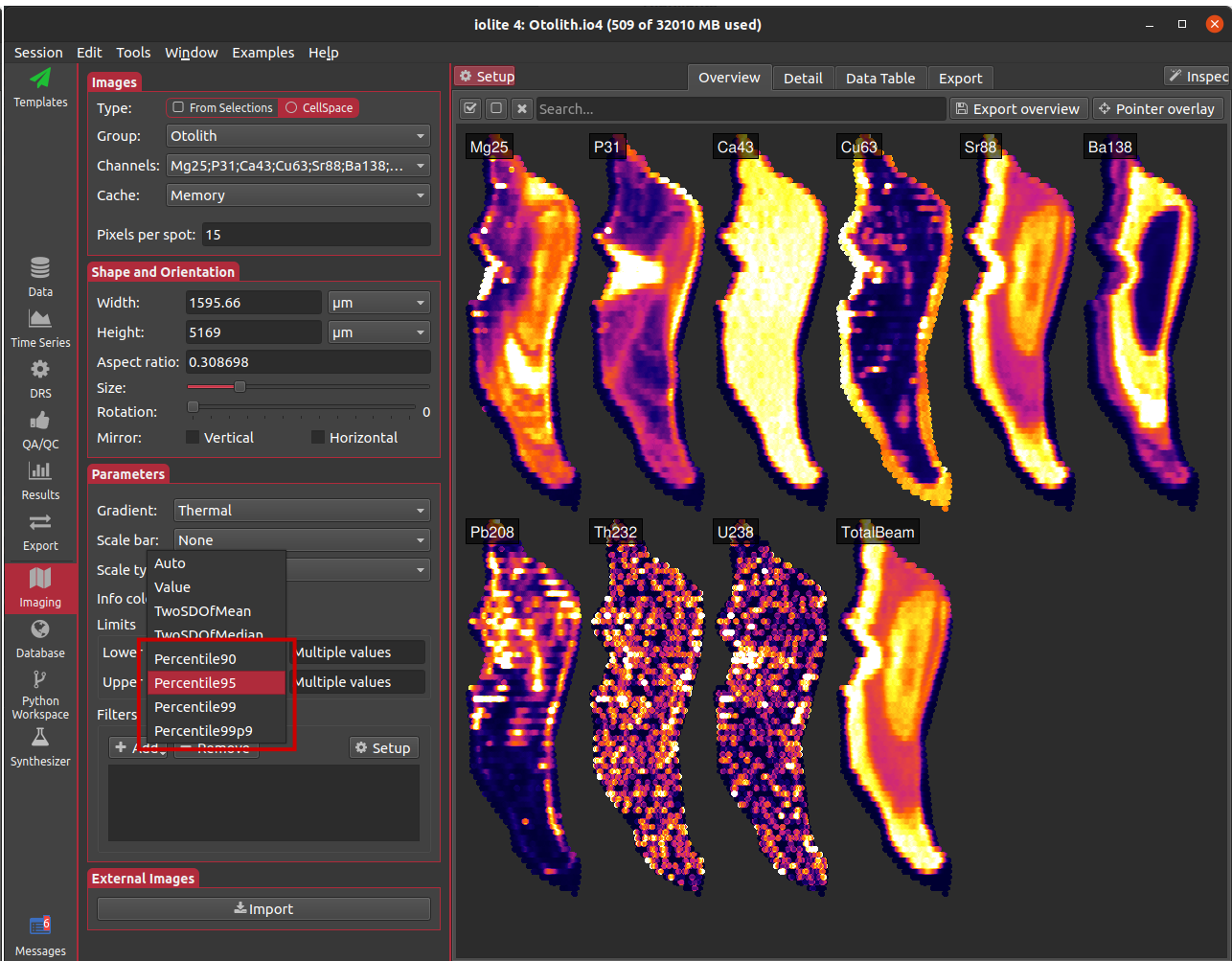
Capturing video from the imaging 3D view
When contrasting two images using iolite’s 3D imaging view, sometimes it is necessary to rotate, pan, and or zoom around the image to see different features of interest. Previously, third party tools were required to capture a video if you wanted to share your observations. Now we have built in a simple tool that allows you to capture mpeg video with the click of a button (the camera) in the 3D imaging tools.

When the button is first clicked, the video file path must be specified. While the view is recording, the button turns red and animates. To stop recording, simply click the button again.
An example recording from our otolith data set is shown below.
Release notes
New features and enhancements:
- Added the ability to zoom when aligning external images
- Added an option to display limits of detection below the results/summary stats plot (for channels that have LODs calculated, e.g. trace elements/ppm channels)
- Modified the behaviour of the Hf isotopes DRS Yb optimization such that a spline can be used rather than the group mean
- Added simple mpeg video capture to the imaging detail/3d view
- Added keyboard shortcuts to python console
- Improved processing templates python interface
- Added option for imaging datatable to display SE or SD
- Added a zip importer that can import data files stored in zip archives
- Added U-Pb downhole fit parameters to settings that are stored in session
- Added some percentile based options for image auto scaling
- Added metadata as an option in results/data table
Recent fixes:
- Fixed an issue with VizualAge and U-Pb QAQC modules where error ellipses were not drawn
- Removed splines from time series view legend
- Fixed an issue with external image alignment dialog when manually aligning images by dragging
- Fixed an issue with external image alignment dialog when no coord file is found
- Fixed some problems with the ‘Trace Elements Next’ DRS when problematic data are encountered
- Fixed selection label header when exporting from results/data table
- Fixed an issue with the concordia age QAQC module causing it fail with an acceptable age
- Fixed an issue where closed QAQC modules would be stored in the session
- Fixed several issues with the beta ‘Trace Elements’ Next data reduction scheme
Click here to discuss.